-Search query
-Search result
Showing all 41 items for (author: mace & k)

EMDB-41265: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (parallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

EMDB-41266: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (antiparallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

PDB-8thi: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (parallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

PDB-8thj: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (antiparallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

EMDB-15775: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Photobacterium profundum in a nanodisc
Method: single particle / : Davies JS, North RA, Dobson RCJ

PDB-8b01: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Photobacterium profundum in a nanodisc
Method: single particle / : Davies JS, North RA, Dobson RCJ

EMDB-13968: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Photobacterium profundum in amphipol
Method: single particle / : North RA, Davies JS, Morado D, Dobson RCJ

PDB-7qha: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Photobacterium profundum in amphipol
Method: single particle / : North RA, Davies JS, Morado D, Dobson RCJ

EMDB-12707: 
O-Layer C14 at 2.58A - Local refinement with C14 symmetry of the O-layer of the outer membrane core complex from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-12708: 
I-Layer C16 at 3.08A - Local refinement with C16 symmetry of I-layer of the outer membrane core complex from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-12709: 
Stalk C1 at 3.71A - Local refinement without symmetry of the Stalk complex from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-12715: 
IMC-Arches C6 at 8.33A - Refinement with C6 symmetry of the Inner Membrane Complex (IMC) with the Arches from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G
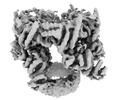
EMDB-12716: 
Trimer of dimers TrwK/VirB4unbound C1 at 4.14A - Refinement without symmetry of TrwK/VirB4unbound trimer of dimers complex (with Hcp1) from the R388 type IV secretion system.
Method: single particle / : Vadakkepat AK, Mace K, Lukoyanova N, Waksman G

EMDB-12717: 
Dimer TrwK/VirB4unbound C1 at 3.49A - Local refinement without symmetry of the TrwK/VirB4unbound dimer complex from the R388 type IV secretion system.
Method: single particle / : Vadakkepat AK, Mace K, Lukoyanova N, Waksman G

EMDB-12933: 
IMC protomer C1 at 3.75A - Local refinement without symmetry of the inner membrane complex (IMC) protomer from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-13765: 
OMCC C1 at 3.28A - Refinement without symmetry of the outer membrane core complex (O- and I-layer) from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Waksman G, Lukoyanova N

EMDB-13766: 
Ab Initio model for IMC-Arches-Stalk from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Waksman G, Lukoyanova N

EMDB-13767: 
IMC-Arches-Stalk C1 at 6.18A - Refinement of the IMC, Arches and Stalk complex without symmetry from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-13768: 
Stalk C5 at 3.28A - Local refinement with C5 symmetry of the Stalk complex from the fully-assembled R388 type IV secretion system.
Method: single particle / : Mace K, Vadakkepat AK, Waksman G, Lukoyanova N

PDB-7o3j: 
O-layer structure (TrwH/VirB7, TrwF/VirB9CTD, TrwE/VirB10CTD) of the outer membrane core complex from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

PDB-7o3t: 
I-layer structure (TrwF/VirB9NTD, TrwE/VirB10NTD) of the outer membrane core complex from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

PDB-7o3v: 
Stalk complex structure (TrwJ/VirB5-TrwI/VirB6) from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

PDB-7o41: 
Hexameric composite model of the Inner Membrane Complex (IMC) with the Arches from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

PDB-7o42: 
TrwK/VirB4unbound trimer of dimers complex (with Hcp1) from the R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Vadakkepat AK, Mace K, Lukoyanova N, Waksman G

PDB-7o43: 
TrwK/VirB4unbound dimer complex from R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Vadakkepat AK, Mace K, Lukoyanova N, Waksman G

PDB-7oiu: 
Inner Membrane Complex (IMC) protomer structure (TrwM/VirB3, TrwK/VirB4, TrwG/VirB8tails) from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

PDB-7q1v: 
Arches protomer (trimer of TrwG/VirB8peri) structure from the fully-assembled R388 type IV secretion system determined by cryo-EM.
Method: single particle / : Mace K, Vadakkepat AK, Lukoyanova N, Waksman G

EMDB-13083: 
L. pneumophila Type IV Coupling Complex (T4CC) with density for DotY N-terminal and middle domains
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

EMDB-13858: 
L. pneumophila Type IV Coupling Complex (T4CC) computed from WT particles where DotY/Z is missing (referred to as T4CCWTminusYZ at 6.3A)
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

EMDB-13859: 
L. pneumophila Type IV Coupling Complex (T4CC) computed from a sample where dotY/Z genes were deleted (referred to as T4CCdeltaDotYZ at 15A)
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

PDB-7ovb: 
L. pneumophila Type IV Coupling Complex (T4CC) with density for DotY N-terminal and middle domains
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

EMDB-10350: 
Type IV Coupling Complex (T4CC) from L. pneumophila.
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

PDB-6sz9: 
Type IV Coupling Complex (T4CC) from L. pneumophila.
Method: single particle / : Mace K, Meir A, Lukoyanova N, Waksman G

EMDB-10288: 
structural insights and activating mutations in diverse pathologies define mechanisms of deregulation for phospholipase C gamma enzymes
Method: single particle / : Liu Y, Bunney T, Phillips C, Katan M

PDB-6r8n: 
STRUCTURE DETERMINATION OF THE TETRAHEDRAL AMINOPEPTIDASE TET2 FROM P. HORIKOSHII BY USE OF COMBINED SOLID-STATE NMR, SOLUTION-STATE NMR AND EM DATA 4.1 A, FOLLOWED BY REAL_SPACE_REFINEMENT AT 4.1 A
Method: single particle / : Colletier JP, Gauto D, Estrozi L, Favier A, Effantin G, Schoehn G, Boisbouvier J, Schanda P

EMDB-4173: 
A FliPQR complex forms the core of the Salmonella type III secretion system export apparatus.
Method: single particle / : Johnson S, Kuhlen L, Abrusci P, Lea SM

PDB-6f2d: 
A FliPQR complex forms the core of the Salmonella type III secretion system export apparatus.
Method: single particle / : Johnson S, Kuhlen L, Abrusci P, Lea SM

EMDB-4179: 
Combined solid-state NMR, solution-state NMR and EM data for structure determination of the tetrahedral aminopeptidase TET2 from P. horikoshii
Method: single particle / : Gauto DF, Estrozi LF, Schwieters CD

PDB-6f3k: 
Combined solid-state NMR, solution-state NMR and EM data for structure determination of the tetrahedral aminopeptidase TET2 from P. horikoshii
Method: single particle / : Gauto DF, Estrozi LF, Schwieters CD, Effantin G, Macek P, Sounier R, Kerfah R, Sivertsen AC, Colletier JP, Boisbouvier J, Schoehn G, Favier A, Schanda P
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
